PCR RFLP
Two tests are described. Both are based on fruit fly-specific PCR of the rDNA ITS region. Differences in method details reflect subtle differences in the standard operating procedures of the original laboratories.
Test 1
Test 1, developed by McKenzie et al. (2004), utilises a 0.6 to 1.2 kb PCR fragment of the ITS1 only. See diagram below and PCR RFLP test 1 for methods.
As the ITS1 can be variable in length, for some species the size of the PCR fragment can be an additional diagnostic character. Separate aliquots of the PCR amplicon are digested with each of up to six different restriction enzymes to produce a species-specific RFLP profile.
This number may be reduced if the potential species could be narrowed down by knowledge of likely origin and/or host, and by reference to the table Analysis of RFLP products from ITS1 fragments from fruit flies, which illustrates which enzymes distinguish which species. This test can be used for identification of 26 fruit fly species (see Diagnostic methods used to identify fruit flies).
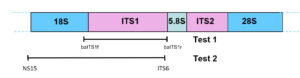
Part of the ribosomal RNA operon with the location of primer positions for Tests 1 and 2
Test 2
Test 2, originally developed by Armstrong and Cameron (1998), is similar to Test 1 but amplifies a larger 1.5-1.8 kb DNA fragment, encompassing the 18S and the ITS1 gene regions. Because of this, the variability in PCR fragment length is not so discernible and is therefore not useful as an additional character. See diagram and PCR RFLP test 2 for methods.
Various combinations of three or four out of 10 restriction enzymes are recommended to produce a species-specific RFLP profile. The specific suite will depend on the likely species identity if it can be narrowed down by likely country of origin or the fruit.
This test can be used for identification of 33 fruit fly species (Diagnostic methods used to identify fruit flies); details are available in Armstrong and Cameron (1998). In the specific circumstance to distinguish just Bactrocera tryoni complex from Ceratitis capitata the enzymes AluI, DdeI, RsaI and SspI are diagnostic.
Species targets
Test 1 can distinguish 25 of 27 target species within the current list of 59 species; all four species within the Bactrocera tryoni group (including B. tryoni, B. neohumeralis, B. melas and B. aquilonis) cannot be distinguished from each other.
Test 2 can distinguish 24 of 33 target species within the current list of species; nine species form four species groups.
Test 2a distinguishes B. tryoni complex from Ceratitis capitata only.
All tests should be considered in terms of the host fruit (for immature life stage samples) and likely country/place of origin to:
- narrow the possible species at the outset for choice of restriction enzymes
- to check the results for reliability.
Host records for the target taxa may assist in the elimination of possible non-target species.
Choice of RFLP test
Test 1 is more rapid, the RFLP patterns less complex to interpret and the variation in size of the ITS1 region easier to detect than Test 2. Therefore this test is recommended when the potential species is within that listed for this test.
Test 2 takes slightly longer, the RFLP patterns are sometimes more complex to interpret and the variation in size of the ITS1 region not as easy to detect is in Test 1.
However, a slightly modified version (Semeraro and Malipatil 2005) provided below, is recommended instead of Test 1 specifically for the diagnosis of the high-risk pest species B. tryoni (although indistinguishable from B. neohumeralis and B. aquilonis within the species complex) and Ceratitis capitata.
The original method provided in Armstrong and Cameron (1998) (not included in Fruit Fly Identification Australia) is also recommended for a further 11 species not included in Test 1 but within the now extended species list on this website; the diagnostic data for Test 2 is however provided in Restriction enzyme haplotype chart and Diagnostic restriction patterns.
Potential for misidentification:
- False positives are possible if other less economically important species, not included during development of the tests, have the same RFLP pattern. This scenario is reduced by increasing the number of restriction sites screened by using several restriction enzymes for an identification. It is also unlikely for exotic fruit fly interceptions given the low potential for non-pest species to reach Australia. For Australian methyl eugenol or cue lure trapped specimens, the range of taxonomically close local non-target species has not been included in development of either test. Therefore care is needed in interpretation of the results in light of other information about the sample, e.g. data on host or geographic origin.
- False negatives could arise if a non-conforming RFLP pattern is produced by a population of a species that has an aberrant polymorphism at a diagnostic restriction site. This is unlikely given the range of populations included during the development of these tests. Also, the ITS1 is generally not useful for detecting population-level variation (cf COI barcodes). False negatives could also arise if the restriction fails or fails to go to completion. DNA from a positive control (known species) should therefore be included to detect this.
Download PCR RFLP test 1 methods
Download PCR RFLP test 2 methods
Choice of standards
All analyses should incorporate at least one standard, i.e. DNA from a known species.
This can include one of the anticipated species for identity of the unknown specimen, which is useful as a positive control to help with sizing the restriction fragments of the unknown.
Alternatively, it can be from a completely unrelated species, which is useful to minimise any question of contamination or mix up.
Standards are essential to provide confidence that the PCR and restriction steps are operating correctly.
